Agarose Gel Simulation
MacVector lets you quickly and easily create simulated agarose gels of your sequences cut with any combination of restriction enzymes that you desire. You can use this feature to identify site(s) that will generate unique patterns to differentiate successful clones, or to determine if fragments are sufficiently different in size to separate under particular gel conditions.
Is It Real Or Is It MacVector?
Agarose Gels appear in separate document-based windows (meaning they can be saved and re-opened) with a realistic appearance;
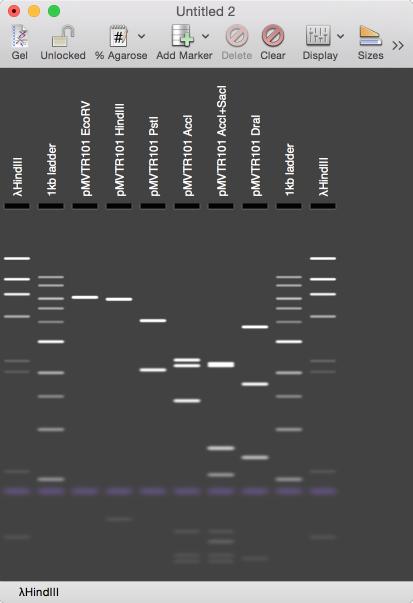

You can select different sets of markers, or define your own. As you mouse-over different bands the status bar updates with details of the fragment(s) under the pointer. The percentage of agarose in the gel and the relative run time can be adjusted as well as the realism of the display. You can optionally view the sizes of the fragments directly on the gel;

When you want to print a gel, you might prefer to see the bands printed as black bands on a white background;
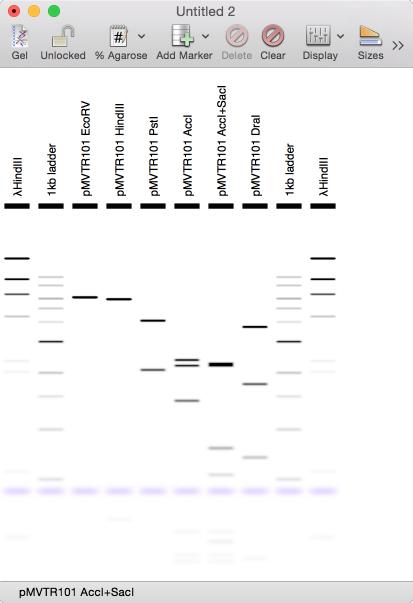

Adding Digests To A Gel
You create a new agarose gel by choosing the File | New | Agarose Gel menu item. Once you have an empty gel (with a default set of size markers that you can control via the Preferences option), you can add lanes to the gel by simply dragging one or more restriction enzymes from the Map tab of a Nucleic Acid Sequence Window, or from the Restriction Enzyme Analysis Results Map tab, and onto the gel. See the animation below;
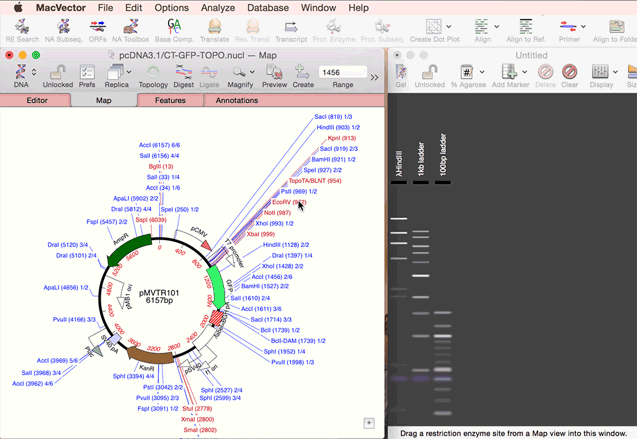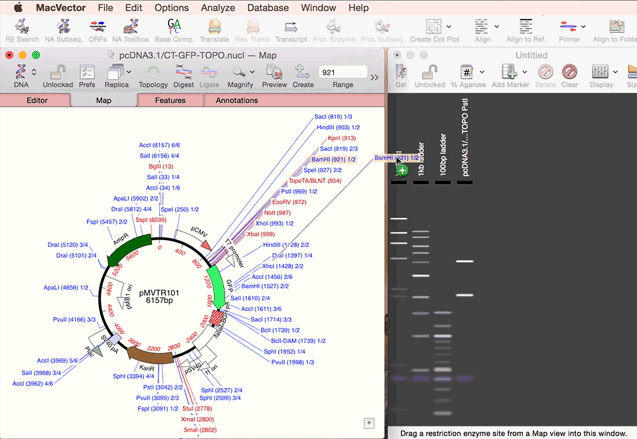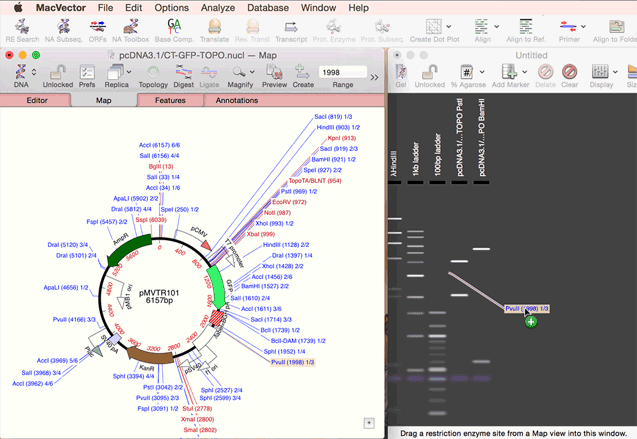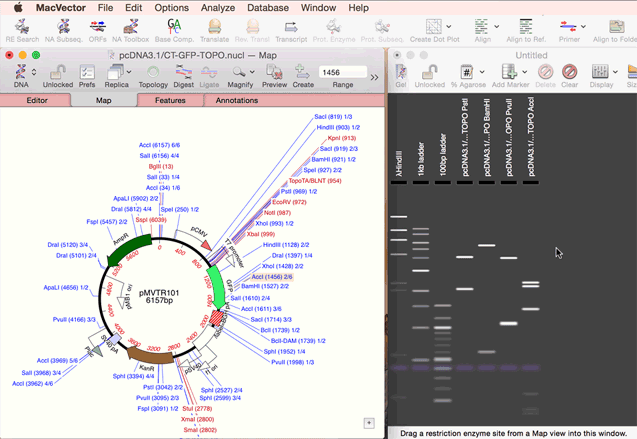
|

2x.png)


2x.png)
